🚨 Paper now online! https://arxiv.org/abs/2111.00595
🚨 Documentation now online! https://mlmed.org/torchxrayvision/
TorchXRayVision
| <img src="https://raw.githubusercontent.com/mlmed/torchxrayvision/master/docs/torchxrayvision-logo.png" width="300px"/> | (🎬 promo video) <br><img src="http://img.youtube.com/vi/Rl7xz0uULGQ/0.jpg" width="400px"/>) |
|---|
What is it?
A library for chest X-ray datasets and models. Including pre-trained models.
TorchXRayVision is an open source software library for working with chest X-ray datasets and deep learning models. It provides a common interface and common pre-processing chain for a wide set of publicly available chest X-ray datasets. In addition, a number of classification and representation learning models with different architectures, trained on different data combinations, are available through the library to serve as baselines or feature extractors.
- In the case of researchers addressing clinical questions it is a waste of time for them to train models from scratch. To address this, TorchXRayVision provides pre-trained models which are trained on large cohorts of data and enables 1) rapid analysis of large datasets 2) feature reuse for few-shot learning.
- In the case of researchers developing algorithms it is important to robustly evaluate models using multiple external datasets. Metadata associated with each dataset can vary greatly which makes it difficult to apply methods to multiple datasets. TorchXRayVision provides access to many datasets in a uniform way so that they can be swapped out with a single line of code. These datasets can also be merged and filtered to construct specific distributional shifts for studying generalization.
Twitter: @torchxrayvision
Getting started
$ pip install torchxrayvision
import torchxrayvision as xrv import skimage, torch, torchvision # Prepare the image: img = skimage.io.imread("16747_3_1.jpg") img = xrv.datasets.normalize(img, 255) # convert 8-bit image to [-1024, 1024] range img = img.mean(2)[None, ...] # Make single color channel transform = torchvision.transforms.Compose([xrv.datasets.XRayCenterCrop(),xrv.datasets.XRayResizer(224)]) img = transform(img) img = torch.from_numpy(img) # Load model and process image model = xrv.models.DenseNet(weights="densenet121-res224-all") outputs = model(img[None,...]) # or model.features(img[None,...]) # Print results dict(zip(model.pathologies,outputs[0].detach().numpy())) {'Atelectasis': 0.32797316, 'Consolidation': 0.42933336, 'Infiltration': 0.5316924, 'Pneumothorax': 0.28849724, 'Edema': 0.024142697, 'Emphysema': 0.5011832, 'Fibrosis': 0.51887786, 'Effusion': 0.27805611, 'Pneumonia': 0.18569896, 'Pleural_Thickening': 0.24489835, 'Cardiomegaly': 0.3645515, 'Nodule': 0.68982, 'Mass': 0.6392845, 'Hernia': 0.00993878, 'Lung Lesion': 0.011150705, 'Fracture': 0.51916164, 'Lung Opacity': 0.59073937, 'Enlarged Cardiomediastinum': 0.27218717}
A sample script to process images usings pretrained models is process_image.py
$ python3 process_image.py ../tests/00000001_000.png
{'preds': {'Atelectasis': 0.50500506,
'Cardiomegaly': 0.6600903,
'Consolidation': 0.30575264,
'Edema': 0.274184,
'Effusion': 0.4026162,
'Emphysema': 0.5036339,
'Enlarged Cardiomediastinum': 0.40989172,
'Fibrosis': 0.53293407,
'Fracture': 0.32376793,
'Hernia': 0.011924741,
'Infiltration': 0.5154413,
'Lung Lesion': 0.22231922,
'Lung Opacity': 0.2772148,
'Mass': 0.32237658,
'Nodule': 0.5091847,
'Pleural_Thickening': 0.5102617,
'Pneumonia': 0.30947986,
'Pneumothorax': 0.24847917}}
Models (demo notebook)
Specify weights for pretrained models (currently all DenseNet121)
Note: Each pretrained model has 18 outputs. The all model has every output trained. However, for the other weights some targets are not trained and will predict randomly becuase they do not exist in the training dataset. The only valid outputs are listed in the field {dataset}.pathologies on the dataset that corresponds to the weights.
## 224x224 models model = xrv.models.DenseNet(weights="densenet121-res224-all") model = xrv.models.DenseNet(weights="densenet121-res224-rsna") # RSNA Pneumonia Challenge model = xrv.models.DenseNet(weights="densenet121-res224-nih") # NIH chest X-ray8 model = xrv.models.DenseNet(weights="densenet121-res224-pc") # PadChest (University of Alicante) model = xrv.models.DenseNet(weights="densenet121-res224-chex") # CheXpert (Stanford) model = xrv.models.DenseNet(weights="densenet121-res224-mimic_nb") # MIMIC-CXR (MIT) model = xrv.models.DenseNet(weights="densenet121-res224-mimic_ch") # MIMIC-CXR (MIT) # 512x512 models model = xrv.models.ResNet(weights="resnet50-res512-all") # DenseNet121 from JF Healthcare for the CheXpert competition model = xrv.baseline_models.jfhealthcare.DenseNet() # Official Stanford CheXpert model model = xrv.baseline_models.chexpert.DenseNet(weights_zip="chexpert_weights.zip") # Emory HITI lab race prediction model model = xrv.baseline_models.emory_hiti.RaceModel() model.targets -> ["Asian", "Black", "White"] # Riken age prediction model model = xrv.baseline_models.riken.AgeModel()
Benchmarks of the modes are here: BENCHMARKS.md and the performance of some of the models can be seen in this paper arxiv.org/abs/2002.02497.
Autoencoders
You can also load a pre-trained autoencoder that is trained on the PadChest, NIH, CheXpert, and MIMIC datasets.
ae = xrv.autoencoders.ResNetAE(weights="101-elastic") z = ae.encode(image) image2 = ae.decode(z)
Segmentation
You can load pretrained anatomical segmentation models. Demo Notebook
seg_model = xrv.baseline_models.chestx_det.PSPNet() output = seg_model(image) output.shape # [1, 14, 512, 512] seg_model.targets # ['Left Clavicle', 'Right Clavicle', 'Left Scapula', 'Right Scapula', # 'Left Lung', 'Right Lung', 'Left Hilus Pulmonis', 'Right Hilus Pulmonis', # 'Heart', 'Aorta', 'Facies Diaphragmatica', 'Mediastinum', 'Weasand', 'Spine']

Datasets
View docstrings for more detail on each dataset and Demo notebook and Example loading script
transform = torchvision.transforms.Compose([xrv.datasets.XRayCenterCrop(), xrv.datasets.XRayResizer(224)]) # RSNA Pneumonia Detection Challenge. https://pubs.rsna.org/doi/full/10.1148/ryai.2019180041 d_kaggle = xrv.datasets.RSNA_Pneumonia_Dataset(imgpath="path to stage_2_train_images_jpg", transform=transform) # CheXpert: A Large Chest Radiograph Dataset with Uncertainty Labels and Expert Comparison. https://arxiv.org/abs/1901.07031 d_chex = xrv.datasets.CheX_Dataset(imgpath="path to CheXpert-v1.0-small", csvpath="path to CheXpert-v1.0-small/train.csv", transform=transform) # National Institutes of Health ChestX-ray8 dataset. https://arxiv.org/abs/1705.02315 d_nih = xrv.datasets.NIH_Dataset(imgpath="path to NIH images") # A relabelling of a subset of NIH images from: https://pubs.rsna.org/doi/10.1148/radiol.2019191293 d_nih2 = xrv.datasets.NIH_Google_Dataset(imgpath="path to NIH images") # PadChest: A large chest x-ray image dataset with multi-label annotated reports. https://arxiv.org/abs/1901.07441 d_pc = xrv.datasets.PC_Dataset(imgpath="path to image folder") # COVID-19 Image Data Collection. https://arxiv.org/abs/2006.11988 d_covid19 = xrv.datasets.COVID19_Dataset() # specify imgpath and csvpath for the dataset # SIIM Pneumothorax Dataset. https://www.kaggle.com/c/siim-acr-pneumothorax-segmentation d_siim = xrv.datasets.SIIM_Pneumothorax_Dataset(imgpath="dicom-images-train/", csvpath="train-rle.csv") # VinDr-CXR: An open dataset of chest X-rays with radiologist's annotations. https://arxiv.org/abs/2012.15029 d_vin = xrv.datasets.VinBrain_Dataset(imgpath=".../train", csvpath=".../train.csv") # National Library of Medicine Tuberculosis Datasets. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4256233/ d_nlmtb = xrv.datasets.NLMTB_Dataset(imgpath="path to MontgomerySet or ChinaSet_AllFiles")
Dataset fields
Each dataset contains a number of fields. These fields are maintained when xrv.datasets.Subset_Dataset and xrv.datasets.Merge_Dataset are used.
-
.pathologiesThis field is a list of the pathologies contained in this dataset that will be contained in the.labelsfield ]. -
.labelsThis field contains a 1,0, or NaN for each label defined in.pathologies. -
.csvThis field is a pandas DataFrame of the metadata csv file that comes with the data. Each row aligns with the elements of the dataset so indexing using.ilocwill work.
If possible, each dataset's .csv will have some common fields of the csv. These will be aligned when The list is as follows:
-
csv.patientidA unique id that will uniqely identify samples in this dataset -
csv.offset_day_intAn integer time offset for the image in the unit of days. This is expected to be for relative times and has no absolute meaning although for some datasets it is the epoch time. -
csv.age_yearsThe age of the patient in years. -
csv.sex_maleIf the patient is male -
csv.sex_femaleIf the patient is female
Dataset tools
relabel_dataset will align labels to have the same order as the pathologies argument.
xrv.datasets.relabel_dataset(xrv.datasets.default_pathologies , d_nih) # has side effects
specify a subset of views (demo notebook)
d_kaggle = xrv.datasets.RSNA_Pneumonia_Dataset(imgpath="...", views=["PA","AP","AP Supine"])
specify only 1 image per patient
d_kaggle = xrv.datasets.RSNA_Pneumonia_Dataset(imgpath="...", unique_patients=True)
obtain summary statistics per dataset
d_chex = xrv.datasets.CheX_Dataset(imgpath="CheXpert-v1.0-small", csvpath="CheXpert-v1.0-small/train.csv", views=["PA","AP"], unique_patients=False) CheX_Dataset num_samples=191010 views=['PA', 'AP'] {'Atelectasis': {0.0: 17621, 1.0: 29718}, 'Cardiomegaly': {0.0: 22645, 1.0: 23384}, 'Consolidation': {0.0: 30463, 1.0: 12982}, 'Edema': {0.0: 29449, 1.0: 49674}, 'Effusion': {0.0: 34376, 1.0: 76894}, 'Enlarged Cardiomediastinum': {0.0: 26527, 1.0: 9186}, 'Fracture': {0.0: 18111, 1.0: 7434}, 'Lung Lesion': {0.0: 17523, 1.0: 7040}, 'Lung Opacity': {0.0: 20165, 1.0: 94207}, 'Pleural Other': {0.0: 17166, 1.0: 2503}, 'Pneumonia': {0.0: 18105, 1.0: 4674}, 'Pneumothorax': {0.0: 54165, 1.0: 17693}, 'Support Devices': {0.0: 21757, 1.0: 99747}}
Pathology masks (demo notebook)
Masks are available in the following datasets:
xrv.datasets.RSNA_Pneumonia_Dataset() # for Lung Opacity xrv.datasets.SIIM_Pneumothorax_Dataset() # for Pneumothorax xrv.datasets.NIH_Dataset() # for Cardiomegaly, Mass, Effusion, ...
Example usage:
d_rsna = xrv.datasets.RSNA_Pneumonia_Dataset(imgpath="stage_2_train_images_jpg", views=["PA","AP"], pathology_masks=True) # The has_masks column will let you know if any masks exist for that sample d_rsna.csv.has_masks.value_counts() False 20672 True 6012 # Each sample will have a pathology_masks dictionary where the index # of each pathology will correspond to a mask of that pathology (if it exists). # There may be more than one mask per sample. But only one per pathology. sample["pathology_masks"][d_rsna.pathologies.index("Lung Opacity")]

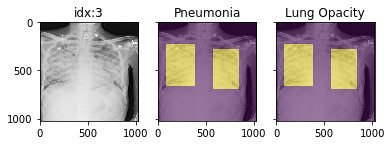
it also works with data_augmentation if you pass in data_aug=data_transforms to the dataloader. The random seed is matched to align calls for the image and the mask.
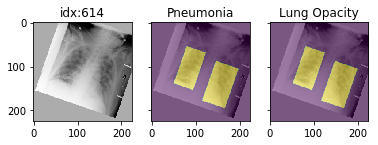

Distribution shift tools (demo notebook)
The class xrv.datasets.CovariateDataset takes two datasets and two
arrays representing the labels. The samples will be returned with the
desired ratio of images from each site. The goal here is to simulate
a covariate shift to make a model focus on an incorrect feature. Then
the shift can be reversed in the validation data causing a catastrophic
failure in generalization performance.
ratio=0.0 means images from d1 will have a positive label ratio=0.5 means images from d1 will have half of the positive labels ratio=1.0 means images from d1 will have no positive label
With any ratio the number of samples returned will be the same.
d = xrv.datasets.CovariateDataset(d1 = # dataset1 with a specific condition d1_target = #target label to predict, d2 = # dataset2 with
编辑推荐精选


音述AI
全球首个AI音乐社区
音述AI是全球首个AI音乐社区,致力让每个人都能用音乐表达自我。音述AI提供零门槛AI创作工具,独创GETI法则帮助用户精准定义音乐风格,AI润色功能支持自动优化作品质感。音述AI支持交流讨论、二次创作与价值变现。针对中文用户的语言习惯与文化背景进行专门优化,支持国风融合、C-pop等本土音乐标签,让技术更好地承载人文表达。


QoderWork
阿里Qoder团队推出的桌面端AI智能体
QoderWork 是阿里推出的本地优先桌面 AI 智能体,适配 macOS14+/Windows10+,以自然语言交互实现文件管理、数据分析、AI 视觉生成、浏览器自动化等办公任务,自主拆解执行复杂工作流,数据本地运行零上传,技能市场可无限扩展,是高效的 Agentic 生产力办公助手。


lynote.ai
一站式搞定所有学习需求
不再被海量信息淹没,开始真正理解知识。Lynote 可摘要 YouTube 视频、PDF、文章等内容。即时创建笔记,检测 AI 内容并下载资料,将您的学习效率提升 10 倍。


AniShort
为AI短剧协作而生
专为AI短剧协作而生的AniShort正式发布,深度重构AI短剧全流程生产模式,整合创意策划、制作执行、实时协作、在线审片、资产复用等全链路功能,独创无限画布、双轨并行工业化工作流与Ani智能体助手,集成多款主流AI大模型,破解素材零散、版本混乱、沟通低效等行业痛点,助力3人团队效率提升800%,打造标准化、可追溯的AI短剧量产体系,是AI短剧团队协同创作、提升制作效率的核心工具。


seedancetwo2.0
能听懂你表达的视频模型
Seedance two是基于seedance2.0的中国大模型,支持图像、视频、音频、文本四种模态输入,表达方式更丰富,生成也更可控。


nano-banana纳米香蕉中文站
国内直接访问,限时3折
输入简单文字,生成想要的图片,纳米香蕉中文站基于 Google 模型的 AI 图片生成网站,支持文字生图、图生图。官网价格限时3折活动


扣子-AI办公
职场AI,就用扣子
AI办公助手,复杂任务高效处理。办公效率低?扣子空间AI助手支持播客生成、PPT制作、网页开发及报告写作,覆盖科研、商业、舆情等领域的专家Agent 7x24小时响应,生活工作无缝切换,提升50%效率!


堆友
多风格AI绘画神器
堆友平台由阿里巴巴设计团队创建,作为一款AI驱动的设计工具,专为设计师提供一站式增长服务。功能覆盖海量3D素材、AI绘画、实时渲染以及专业抠图,显著提升设计品质和效率。平台不仅提供工具,还是一个促进创意交流和个人发展的空间,界面友好,适合所有级别的设计师和创意工作者。


码上飞
零代码AI应用开发平台
零代码AI应用开发平台,用户只需一句话简单描述需求,AI能自动生成小程序、APP或H5网页应用,无需编写代码。

Vora
免费创建高清无水印Sora视频
Vora是一个免费创建高清无水印Sora视频的AI工具
推荐工具精选
AI云服务特惠
懂AI专属折扣关注微信公众号
最新AI工具、AI资讯
独家AI资源、AI项目落地

微信扫一扫关注公众号






